Accurately identifying and annotating cell types is a crucial step in single-cell RNA-Seq analysis. Often, this task requires experience and specialized domain-level knowledge. This post describes how NEBION proceeds with establishing cell-type references that allow a precise and accurate annotation of single-cell RNA-Seq studies.
Choose the right data
For a given body system, NEBION curators first scout public sources to identify high-quality single-cell studies having a broad range of cell types and a sufficient number of replicates (subjects). Based on the corresponding publications, we then select a core set of studies having the broadest possible number of cell types and good coverage.
Identify cell types
All data is then processed and analyzed by experts to identify all known and novel cell types, using both published and proprietary markers.
Harmonize across studies
A key aim of NEBION is to make all curated data sets comparable across studies from different origins. This means that cell types need to be well characterized using a common set of vocabularies (cell-type ontology). All studies are then cross-compared and harmonized such as to define a core set of coordinates and markers for every cell type. A cell-type reference is then generated and used subsequently for the annotation of new studies.
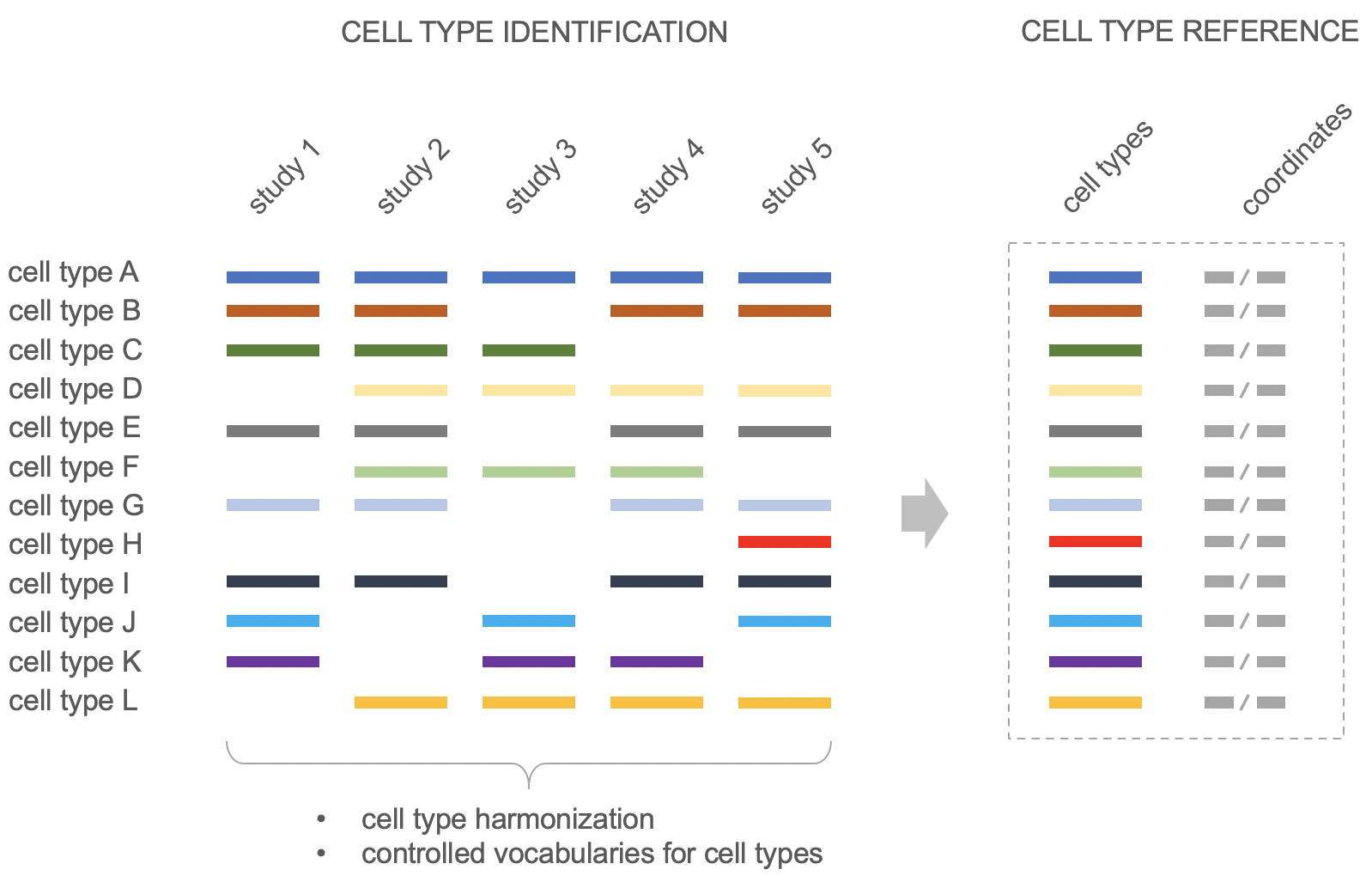 Figure: Schematic representation of the establishment of cell-type references at NEBION. All studies from the same organ are fully harmonized, using a common set of controlled vocabularies and a common cell-type reference. This allows users to easily perform cross-study analyses in GENEVESTIGATOR®.
Figure: Schematic representation of the establishment of cell-type references at NEBION. All studies from the same organ are fully harmonized, using a common set of controlled vocabularies and a common cell-type reference. This allows users to easily perform cross-study analyses in GENEVESTIGATOR®.
Accurately analyze!
Having a robust and well tested cell-type reference significantly speeds up the analysis of single-cell RNA-Seq data and delivers more accurate results. NEBION offers cell-type references for various body systems for licensing to commercial customers. In parallel, public single-cell studies curated by NEBION get integrated into GENEVESTIGATOR® for seamless compendium-wide and cell-type level analysis. To learn more about our cell-type references and licensing conditions, contact us at sales@nebion.com.